We are excited to share new research from our collaboration with the Phillips Lab at UCSF, now on bioRxiv!
How do antibodies evolve to recognize emerging SARS-CoV-2 variants without losing the biophysical properties needed to function? By characterizing all possible evolutionary intermediates of a human antibody lineage, the Phillips Lab found that only a handful of mutational paths can satisfy these competing demands — and that those paths acquire mutations in a very specific order.
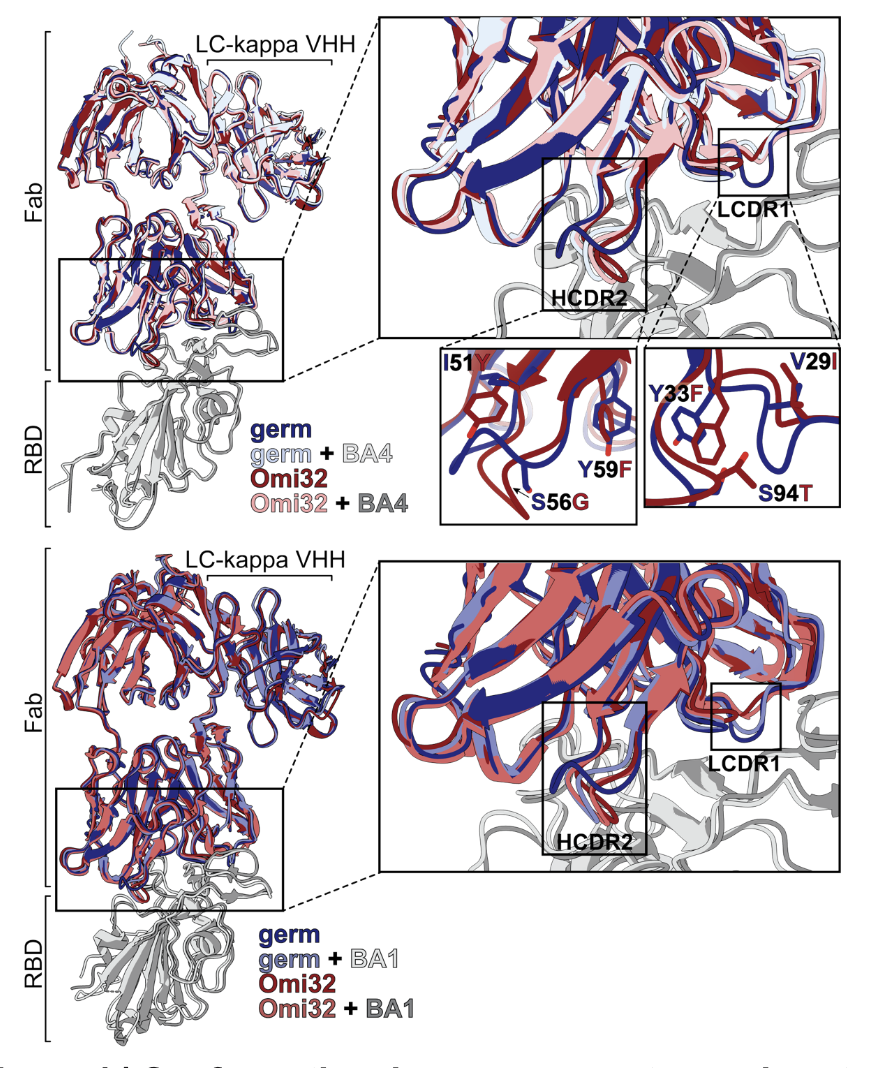
NanoImaging Services performed all cryo-EM structure determination for this project, enabling epitope mapping of both the ancestral and mature antibodies. These structures revealed a conformational rearrangement that relieves steric clashes — unlocking mutational paths that would otherwise be evolutionarily inaccessible.
Those of you who joined our webinar, "Structural Heterogeneity in Focus: Preparing for Cryo-EM Success", may recognize this data, which was used to highlight how cryo-EM can be used for epitope mapping of structurally heterogeneous samples.
Great science doesn't happen without great partners. Thank you to the Phillips Lab for a fantastic collaboration!
We are excited to share new research from our collaboration with the Phillips Lab at UCSF, now on bioRxiv!
How do antibodies evolve to recognize emerging SARS-CoV-2 variants without losing the biophysical properties needed to function? By characterizing all possible evolutionary intermediates of a human antibody lineage, the Phillips Lab found that only a handful of mutational paths can satisfy these competing demands — and that those paths acquire mutations in a very specific order.
NanoImaging Services performed all cryo-EM structure determination for this project, enabling epitope mapping of both the ancestral and mature antibodies. These structures revealed a conformational rearrangement that relieves steric clashes — unlocking mutational paths that would otherwise be evolutionarily inaccessible.
Those of you who joined our webinar, "Structural Heterogeneity in Focus: Preparing for Cryo-EM Success", may recognize this data, which was used to highlight how cryo-EM can be used for epitope mapping of structurally heterogeneous samples.
Great science doesn't happen without great partners. Thank you to the Phillips Lab for a fantastic collaboration!
Tharp, C. R., Catalano, C., Khalifeh, A., Ghaffari-Kashani, S., Huang, R., Kang, G., Scapin, G., & Phillips, A. M. (2026). Biophysical trade-offs in antibody evolution are resolved by conformation-mediated epistasis. bioRxiv. https://doi.org/10.64898/2026.03.12.711465